New paper by Roland K.O. Sigel in PNAS
Roland Sigel published together with the group of David Rueda Imperial College a paper in PNAS (Proceedings of the National Academy of Sciences of the U.S.A.) entitled Optimal molecular crowding accelerates group II intron folding and maximizes catalysis.
Biological processes take place in living cells and have adapted to environmental conditions such as temperature, salt concentration, and high-density cellular contents. However, our molecular understanding of most biological processes relies on in vitro experiments under nonphysiological, highly diluted, and high-salt conditions. To overcome this, we mimic the cellular environment by introducing crowding agents. To understand how such conditions affect large-ribozyme folding and function, we studied a group IIB intron ribozyme under crowded, low-salt conditions. The data show how crowded environments enhance the activity of such large ribozymes, even in low-salt concentrations. Interestingly, we find that “optimal” crowding yields maximum activity, and we are the first reporting that beyond optimal conditions dense crowding becomes detrimental for ribozymes activity.
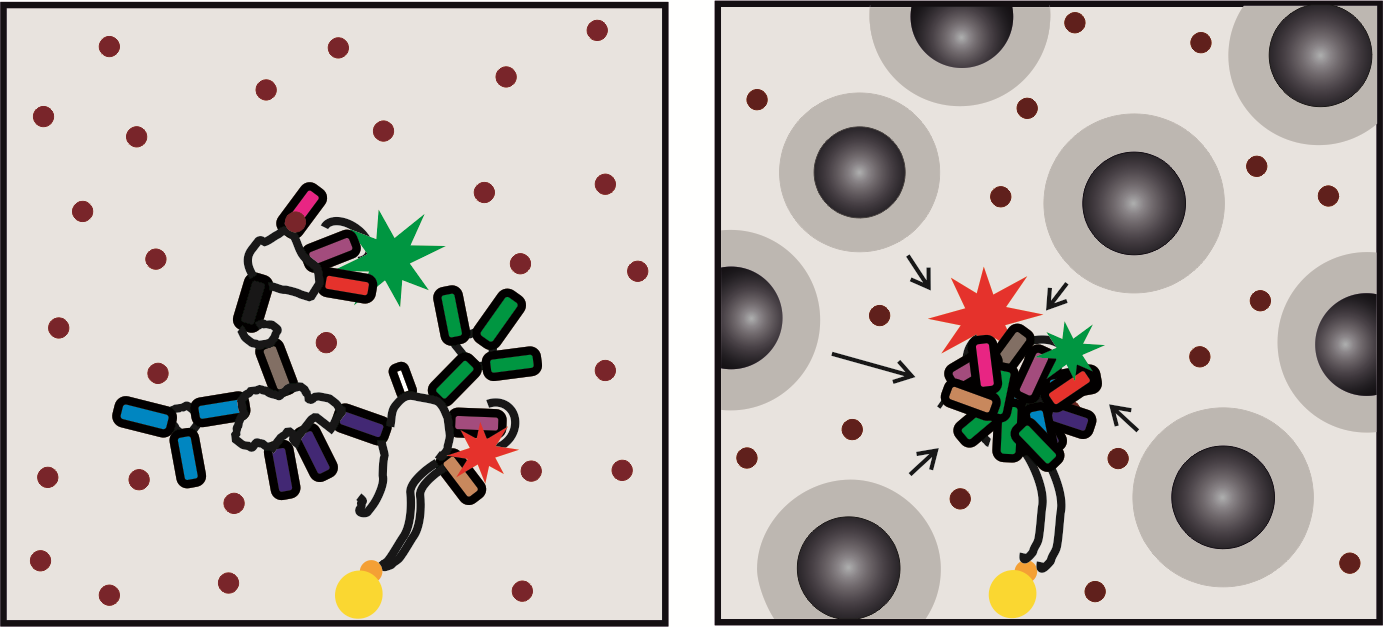
TOC graph:
Reference:
Optimal molecular crowding accelerates group II intron folding and maximizes catalysis
Bishnu P. Paudel*, Erica Fiorini*, Richard Börner*, Roland K. O. Sigel+, David S. Rueda+
Proceedings of the National Academy of Sciences Nov 2018, 201806685; DOI: 10.1073/pnas.1806685115
Richard Börner