Chemical Biology
RNA Chemistry
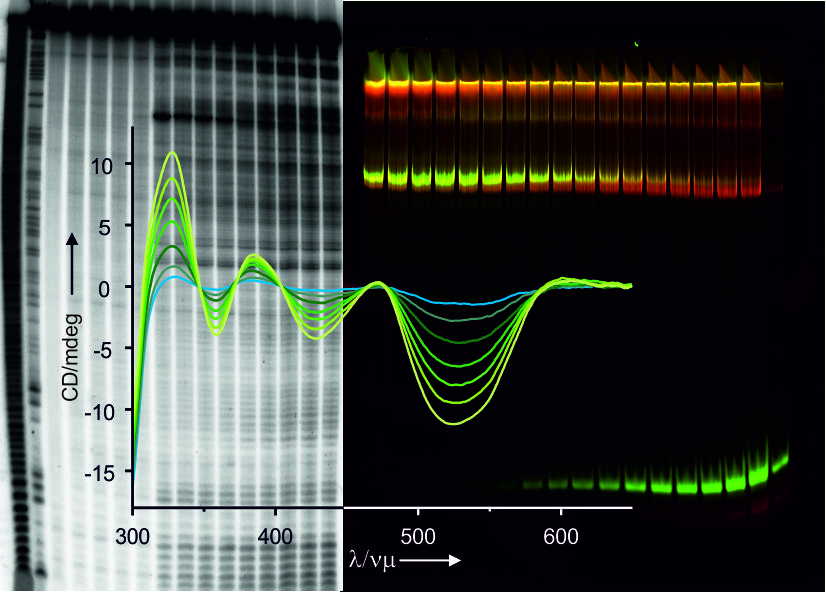
To investigate structural characteristics of RNA molecules, as well as to monitor their interactions with metal ions or metal complexes, we use a number of spectroscopic and biochemical techniques. Spectroscopic methods comprise UV/VIS absorption, including RNA thermal melting, and fluorescence emission, as well as dynamic light scattering (DLS) and circular dichroism (CD). Biochemical methods include chemical and enzymatic footprinting assays designed to determine specific metal ion binding sites, local structural changes, large structural rearrangements, and RNA interactions with other molecules such as proteins. These assays are combined with polyacrylamide gel electrophoresis (PAGE) and include, beside others, in-line probing, terbium cleavage and OH-radical cleavage. Generally, our footprinting assays are performed with 32P 3’- or 5’-end radioactively labeled RNA using phosphoimaging for visualization [1-4]. In addition, we use fluorescently end-labeled RNA or we take advantage form fluorescently labeled DNA and PNA oligonucleotides as hybridization probes [5] using fluorescence gel imaging. For smFRET studies we have further co-developed a method for site-specific internal labeling of large RNAs, such as group II introns and riboswitches [6,7].
Literature
[1] Michelle F. Schaffer, Pallavi K. Choudhary, Roland K.O. Sigel*, Methods Enzymol., 2015, 549, 467-488.
doi:10.1016/B978-0-12-801122-5.00020-9
[2] Pallavi K. Choudhary, Sofia Gallo, Roland K.O. Sigel*, Methods Mol. Biol., 2014, 1086, 143-158.
doi:10.1007/978-1-62703-667-2_8
[3] Michèle C. Erat, Roland K.O. Sigel*, Met. Ions Life Sci., 2011, 9, 37-100.
doi:10.1039/9781849732512-00037
[4] Maria Pechlaner, Roland K.O. Sigel*, Met. Ions Life Sci., 2012, 10, 1-42.
doi:10.1007/978-94-007-2172-2_1
[5] Anita G. Schmitz, Susann Zelger-Paulus, Gilles Gasser*, Roland K. O. Sigel*, ChemBioChem, 2015, 16, 1302-1306.
doi:10.1002/cbic.201500180
[6] David Egloff, Igor A. Oleinich, Sebastian L. B. König, Roland K. O. Sigel, Eva Freisinger*, ACS Chem. Biol., 2016, 11, 2558-2567.
doi:10.1021/acschembio.6b00343
[7] Zhao, Meng; Steffen, Fabio D.; Börner, Richard; Sigel, Roland K. O.; Freisinger, Eva; Nucleic Acids Research 2017, published online.
doi:10.1093/nar/gkx1100